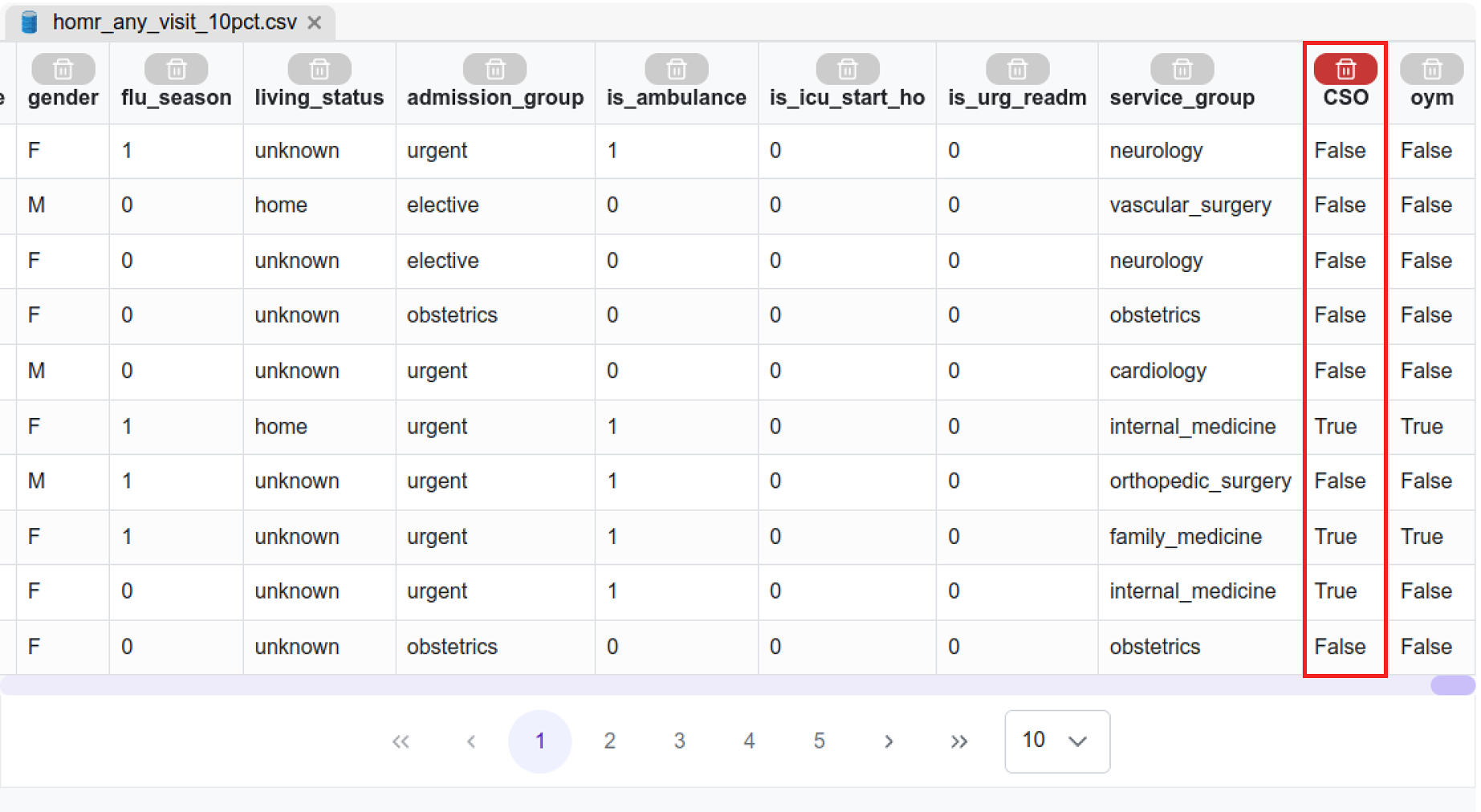
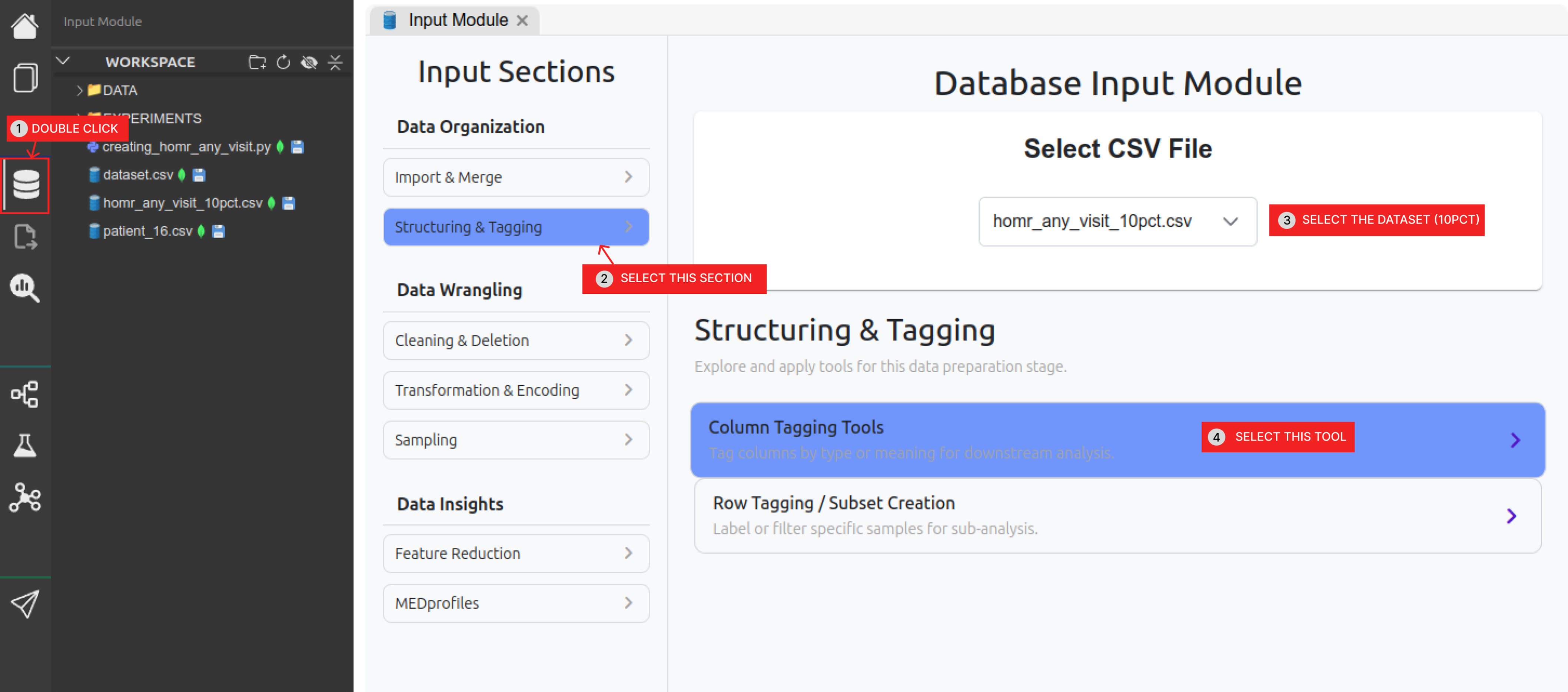
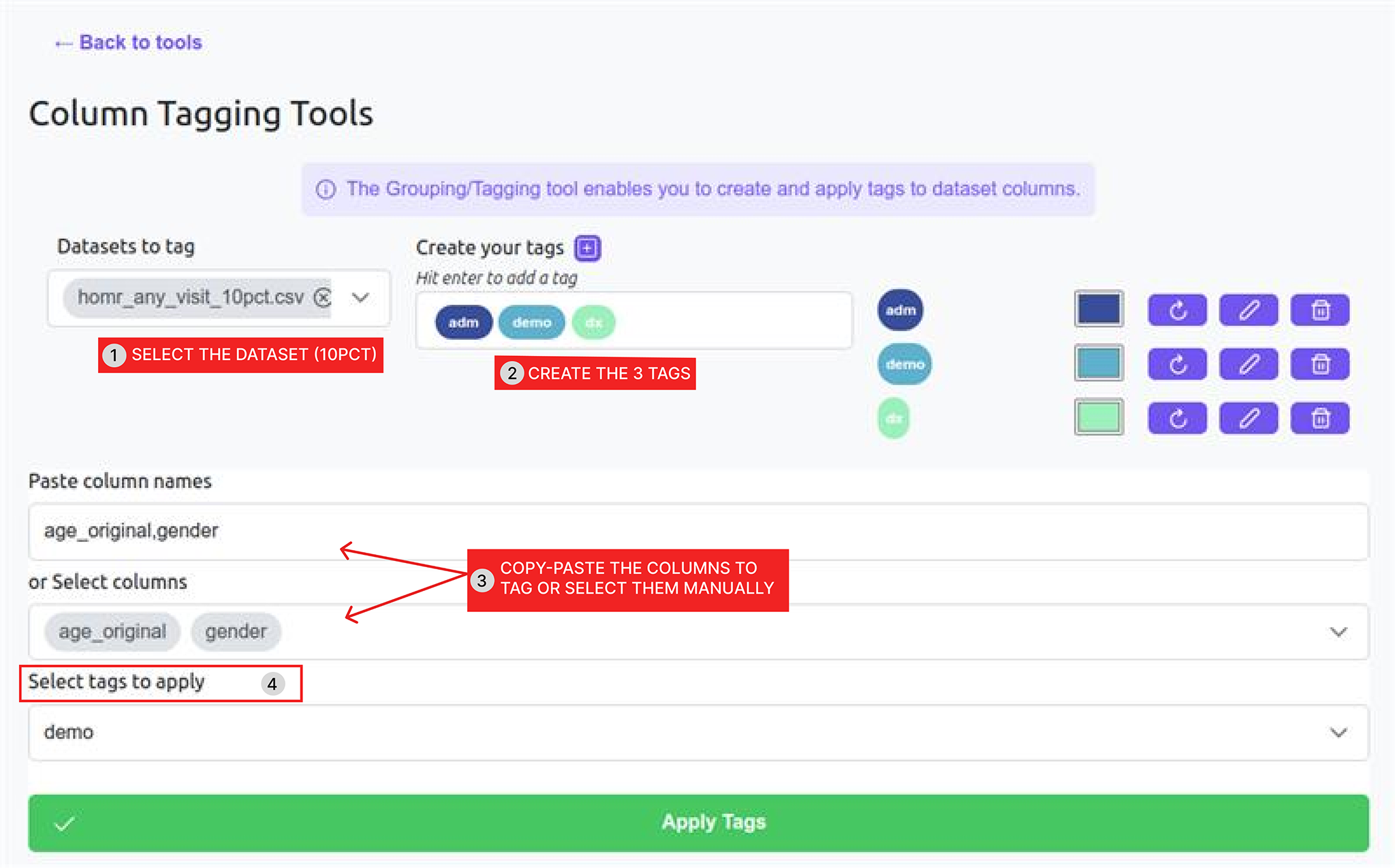
| Dx | Comorbidity diagnoses + Admission diagnoses (the rest of the columns)
Simply copy paste the following code line into the tagging tool: dx_pneumo_adm,dx_obstructive,dx_asthma,dx_bronchiectasis,dx_chronic_resp_failure,dx_acute_resp_failure,dx_ild,dx_home_o2,dx_pseudomonas,dx_pulmonary_hypertension,dx_obesity_hypoventilation,dx_pneumonia_adm,dx_recent_pneumonia,dx_liver_1,dx_liver_2,dx_liver_rf,dx_ascites,dx_anasarca,dx_alcohol,dx_ibd_crohn,dx_recent_abdominal_pain,dx_recent_intestinal_occlusion,dx_recent_gi_bleed,dx_recent_colitis,dx_recent_perforation,dx_renal_1,dx_renal_2,dx_dialysis,dx_recent_interstitial_nephritis,dx_recent_uti,dx_dementia,dx_frailty,dx_denutrition,dx_falls,dx_cachexia,dx_paralysis,dx_cvd,dx_psych,dx_depression,dx_endo_1,dx_endo_2,dx_mi_recent,dx_mi_past,dx_angina_recent,dx_chf,dx_chf_adm,dx_cad,dx_valve,dx_aortic_stenosis,dx_a_fib,dx_recent_chest_pain,dx_pvd,dx_recent_hip_fracture,dx_recent_back_pain,dx_anticoagulation,dx_anemia,dx_recent_anemia,dx_past_pe,dx_recent_pe,dx_orl_cancer,dx_gi_cancer_1,dx_gi_cancer_2,dx_gi_cancer_3,dx_chest_cancer_1,dx_chest_cancer_2,dx_msk_cancer,dx_skin_cancer,dx_breast_cancer,dx_gu_cancer_1,dx_gu_cancer_2,dx_gu_cancer_3,dx_cns_cancer,dx_endocrine_cancer,dx_heme_cancer_1,dx_heme_cancer_2,dx_heme_cancer_3,dx_metastatic_solid_cancer,dx_cancer_ed,dx_chemo_cancer_1,dx_chemo_cancer_2,dx_palliative,dx_transplant,dx_recent_complication,dx_obstetrics,has_dx,adm_abcess,adm_abdominal_pain,adm_acute_leukemia,adm_alcohol,adm_tonsillitis,adm_anemia,adm_aneurism,adm_angina,adm_aortic_aneurism,adm_appendicitis,adm_ards,adm_arrhythmia,adm_arthropathy,adm_ascites,adm_aspiration_pneumonia,adm_asthma,adm_atrial_fibrillation,adm_bariatric,adm_benign_tumor,adm_bi_pan_cytopenia,adm_biliary_colic,adm_bladder_cancer,adm_brain_cancer,adm_brain_hemorrhage,adm_brain_injury,adm_brain_lesion,adm_breast_cancer,adm_bronchiectasis,adm_bronchitis,adm_c_difficile,adm_cancer,adm_carotid_stenosis,adm_cellulitis,adm_chemotherapy,adm_chest_pain,adm_cholangitis,adm_cholecystitis,adm_choledocholithiasis,adm_chronic_leukemia,adm_cirrhosis,adm_colorectal_cancer,adm_colitis,adm_conduction_abnormality,adm_copd,adm_delirium,adm_dementia,adm_diabetes,adm_dialysis,adm_diarrhea,adm_disk_disorder,adm_diverticular_disease,adm_dvt,adm_dysphagia,adm_dyspnea,adm_ear_disorder,adm_electrolytes,adm_endocarditis,adm_eol_care,adm_eps,adm_eye,adm_falls,adm_febrile_neutropenia,adm_fertility,adm_fracture,adm_gastric_cancer,adm_gastritis,adm_enteritis,adm_gi_bleed,adm_guillain_barre,adm_gyn_disorder,adm_heart_failure,adm_hemoptysis,adm_hepatic_failure,adm_hepatitis,adm_hip_fracture,adm_hypertension,adm_infection,adm_inguinal_hernia,adm_intestinal_ischemia,adm_intestinal_polyp,adm_intoxication,adm_joint_prosthesis,adm_liver_cancer,adm_loss_of_autonomy,adm_lower_leg_fracture,adm_lumbar_pelvis_fracture,adm_lung_cancer,adm_lung_mass,adm_lymphoma,adm_melanoma,adm_meningitis,adm_metastasis,adm_mi,adm_mii,adm_multiple_myeloma,adm_oesophageal_cancer,adm_oesophageal_varices,adm_orl_cancer,adm_osteomyelitis,adm_osteoporosis,adm_other_hernia,adm_pancreatic_cancer,adm_pancreatic_mass,adm_pancreatitis,adm_parkinsons,adm_perforation,adm_pericardial_effusion,adm_pericarditis,adm_pleural_effusion,adm_pneumonia,adm_pneumothorax,adm_pregnancy,adm_prolapsus,adm_prostate_cancer,adm_pulmonary_fibrosis,adm_pulmonary_hypertension,adm_pvd,adm_pvd_gangrene,adm_pvd_insufficiency,adm_pvd_ischemia,adm_reanimation,adm_renal_failure,adm_respiratory_failure,adm_seizures,adm_sepsis,adm_severe,adm_shock,adm_spondylopathy,adm_stroke,adm_subarachnoid_hemorrhage,adm_syncope_hypotension,adm_tachycardia,adm_tamponnade,adm_thyroid_cancer,adm_tia,adm_trauma,adm_trigeminal_neuralgia,adm_tumor,adm_urinary_lithiasis,adm_urinary_retention,adm_uro_procedure,adm_uti,adm_valve_prosthesis,adm_valve_regurgitation,adm_valve_stenosis,adm_virus,adm_weight_loss_fatiguedx_pneumo_adm,dx_obstructive,dx_asthma,dx_bronchiectasis,dx_chronic_resp_failure,dx_acute_resp_failure,dx_ild,dx_home_o2,dx_pseudomonas,dx_pulmonary_hypertension,dx_obesity_hypoventilation,dx_pneumonia_adm,dx_recent_pneumonia,dx_liver_1,dx_liver_2,dx_liver_rf,dx_ascites,dx_anasarca,dx_alcohol,dx_ibd_crohn,dx_recent_abdominal_pain,dx_recent_intestinal_occlusion,dx_recent_gi_bleed,dx_recent_colitis,dx_recent_perforation,dx_renal_1,dx_renal_2,dx_dialysis,dx_recent_interstitial_nephritis,dx_recent_uti,dx_dementia,dx_frailty,dx_denutrition,dx_falls,dx_cachexia,dx_paralysis,dx_cvd,dx_psych,dx_depression,dx_endo_1,dx_endo_2,dx_mi_recent,dx_mi_past,dx_angina_recent,dx_chf,dx_chf_adm,dx_cad,dx_valve,dx_aortic_stenosis,dx_a_fib,dx_recent_chest_pain,dx_pvd,dx_recent_hip_fracture,dx_recent_back_pain,dx_anticoagulation,dx_anemia,dx_recent_anemia,dx_past_pe,dx_recent_pe,dx_orl_cancer,dx_gi_cancer_1,dx_gi_cancer_2,dx_gi_cancer_3,dx_chest_cancer_1,dx_chest_cancer_2,dx_msk_cancer,dx_skin_cancer,dx_breast_cancer,dx_gu_cancer_1,dx_gu_cancer_2,dx_gu_cancer_3,dx_cns_cancer,dx_endocrine_cancer,dx_heme_cancer_1,dx_heme_cancer_2,dx_heme_cancer_3,dx_metastatic_solid_cancer,dx_cancer_ed,dx_chemo_cancer_1,dx_chemo_cancer_2,dx_palliative,dx_transplant,dx_recent_complication,dx_obstetrics,has_dx,adm_abcess,adm_abdominal_pain,adm_acute_leukemia,adm_alcohol,adm_tonsillitis,adm_anemia,adm_aneurism,adm_angina,adm_aortic_aneurism,adm_appendicitis,adm_ards,adm_arrhythmia,adm_arthropathy,adm_ascites,adm_aspiration_pneumonia,adm_asthma,adm_atrial_fibrillation,adm_bariatric,adm_benign_tumor,adm_bi_pan_cytopenia,adm_biliary_colic,adm_bladder_cancer,adm_brain_cancer,adm_brain_hemorrhage,adm_brain_injury,adm_brain_lesion,adm_breast_cancer,adm_bronchiectasis,adm_bronchitis,adm_c_difficile,adm_cancer,adm_carotid_stenosis,adm_cellulitis,adm_chemotherapy,adm_chest_pain,adm_cholangitis,adm_cholecystitis,adm_choledocholithiasis,adm_chronic_leukemia,adm_cirrhosis,adm_colorectal_cancer,adm_colitis,adm_conduction_abnormality,adm_copd,adm_delirium,adm_dementia,adm_diabetes,adm_dialysis,adm_diarrhea,adm_disk_disorder,adm_diverticular_disease,adm_dvt,adm_dysphagia,adm_dyspnea,adm_ear_disorder,adm_electrolytes,adm_endocarditis,adm_eol_care,adm_eps,adm_eye,adm_falls,adm_febrile_neutropenia,adm_fertility,adm_fracture,adm_gastric_cancer,adm_gastritis,adm_enteritis,adm_gi_bleed,adm_guillain_barre,adm_gyn_disorder,adm_heart_failure,adm_hemoptysis,adm_hepatic_failure,adm_hepatitis,adm_hip_fracture,adm_hypertension,adm_infection,adm_inguinal_hernia,adm_intestinal_ischemia,adm_intestinal_polyp,adm_intoxication,adm_joint_prosthesis,adm_liver_cancer,adm_loss_of_autonomy,adm_lower_leg_fracture,adm_lumbar_pelvis_fracture,adm_lung_cancer,adm_lung_mass,adm_lymphoma,adm_melanoma,adm_meningitis,adm_metastasis,adm_mi,adm_mii,adm_multiple_myeloma,adm_oesophageal_cancer,adm_oesophageal_varices,adm_orl_cancer,adm_osteomyelitis,adm_osteoporosis,adm_other_hernia,adm_pancreatic_cancer,adm_pancreatic_mass,adm_pancreatitis,adm_parkinsons,adm_perforation,adm_pericardial_effusion,adm_pericarditis,adm_pleural_effusion,adm_pneumonia,adm_pneumothorax,adm_pregnancy,adm_prolapsus,adm_prostate_cancer,adm_pulmonary_fibrosis,adm_pulmonary_hypertension,adm_pvd,adm_pvd_gangrene,adm_pvd_insufficiency,adm_pvd_ischemia,adm_reanimation,adm_renal_failure,adm_respiratory_failure,adm_seizures,adm_sepsis,adm_severe,adm_shock,adm_spondylopathy,adm_stroke,adm_subarachnoid_hemorrhage,adm_syncope_hypotension,adm_tachycardia,adm_tamponnade,adm_thyroid_cancer,adm_tia,adm_trauma,adm_trigeminal_neuralgia,adm_tumor,adm_urinary_lithiasis,adm_urinary_retention,adm_uro_procedure,adm_uti,adm_valve_prosthesis,adm_valve_regurgitation,adm_valve_stenosis,adm_virus,adm_weight_loss_fatigue
| 232 |